GENONE
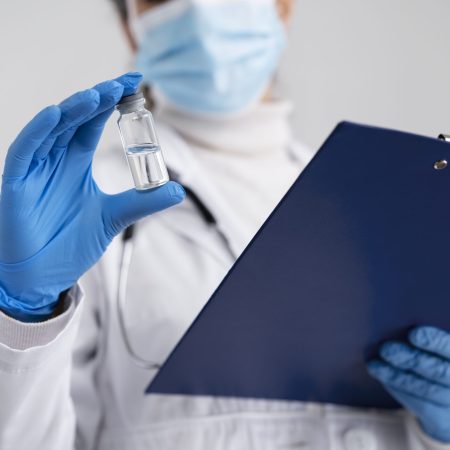
Síntese de Peptídeos
Os peptídeos sintéticos oferecem vantagens para diversas aplicações, indo desde novas drogas até a pesquisa do funcionamento e interação de proteínas.
Os peptídeos sintéticos oferecem vantagens para diversas aplicações, indo desde novas drogas até a pesquisa do funcionamento e interação de proteínas.
-
Recomendação
de Pureza
-
Modificações
Realizadas
-
Cell Penetrating
Peptides
-
Peptídeos
Catalogados
-
Bancos de dados
de Peptídeos
A GenOne Biotechnologies propõe a seguir uma tabela com a pureza dos peptídeos recomendada para os ensaios mais comuns solicitados pelos nossos clientes.
Pureza recomendada
Aplicações
Imunologia Pureza - 70%
- Antígeno para a produção e purificação por afinidade de anticorpos policlonais
- Testes de ELISA
- Peptide arrays
Bioquímica Pureza - 85%
- Mapeamento de epítopos
- Estudos semi-quantitativos de interação enzima-substrato
- Estudos de fosforilação
- Estudos de peptide blocking por Western blotting
- Ensaios in vitro
- Estudos de aderência celular
Alta Pureza Pureza - 95%
- Estudos quantitativos de interação receptor-ligante
- Ensaios quantitativos de inibição e bloqueio competitivos
- Estudos quantitativos de fosforilação
- Estudos quantitativosde proteólise
- Estudos de NMR
- Estudos in vivo
- Geração de curvas de padrão
Industrial Pureza - 98%
- Peptídeos BPF para estudos de drogas
- Peptídeos para a indústria cosmética
- Ensaios clínicos
- Estudos de relação estrutura x atividade (SAR)
Observações:
Quando um peptídeo é entregue com pureza de 95%, isso significa que 95% do conteúdo de peptídeo entregue é composto pelo peptídeo solicitado (o que difere do conteúdo total do peptídeo). Os outros 5% de material peptídico é usualmente composto do que é chamado de “sequências deletadas” que usualmente podem ser co-purificadas com o peptídeo desejado durante a HPLC.
As sequências deletadas são geradas durante a síntese dos peptídeos, quando devido a ineficiência da reação de acoplamento, alguns aminoácidos são omitidos das moléculas sintetizadas. A pureza é determinada por HPLC de fase reversa.
É importante entender a diferença entre conteúdo peptídico e conteúdo total de peptídeos. Os peptídeos secos e entregues na forma de pó, normalmente não possuem somente peptídeos, mas também algumas substâncias como água, solventes adsorvidos, contra-íons e sais. Devido a própria natureza dos peptídeos, dificuldade de solubilidade, haverá quase sempre outros aditivos de forma a permitir solubilidade e estabilidade.
O conteúdo total se refere a esta mistura enquanto o conteúdo peptídico refere-se à fração do seu peptídeo na mistura. O conteúdo peptídico é usualmente 50-80% da massa total de pó enviado, esse conteúdo só é obtido precisamente pela análise do conteúdo de aminoácidos ou quantidade de nitrogênio que pode ser solicitada como um serviço à parte, bem como a remoção de TFA.
Existe uma maneira de se estimar o conteúdo peptídico teórico, assumindo que o único conteúdo não peptídico presente na amostra são os contra-íons. É possível fazer isso dividindo a massa molecular do peptídeo pela soma da massa molecular com o número de contra-íons necessários para neutralizar o peptídeo. O número de contra-íons é multiplicado pela massa molecular do TFA (MW=114).
Por exemplo, um peptídeo com massa molecular de 1500, com N-Terminal livre e uma Arginina, o seu conteúdo líquido é = 1500/(1500+2×114)= 0,87 ou 87%. Na prática, os contra-íons não são os únicos contaminantes, como citado anteriormente há também algumas substâncias como água, solventes adsorvidos e sais.
A seguir mostramos a grande diversidade de modificações que podem ser realizadas ao fazer o seu pedido com a GenOne Biotechnologies, porém caso deseje alguma que não esteja na lista não hesite em solicitar mais informações.
Modificações N-Terminal
Modificações C-Terminal
Modificações diversas
Acetilatação (Ac)
Amidação
Biotinilação, Fluoróforos
Formilação (For)
AMC
Pontes dissulfeto
Ácidos Graxos
Aldeído
Fosforilação
Benzoil (Bz)
Alcool
Conjugação com BSA, KLH, OVA
Benziloxicarbonilação (CBZ)
Éster (OMe, OEt)
PEGlação
Bromoacetil (Br-Ac)
p-Nitroanilina (pNA)
MAPS
Piroglutamil (pGlu) (Pyr)
NHMe, NHEt e NHisopen
Peptídeos Cíclicos
Succinilação (Suc)
tBu
Fluorescência e Quenchers
Tert-Butoxicarbonil (Boc)
TBzl
Aminoácidos Especiais
3-Mercaptopropil (Mpa)
Cisteamida (Cya)
Metil e N-Metil
Os CPPs são capazes de transportar ácidos nucleicos e nanomateriais para o interior das células.
Tabela em breve
Entre em contato conosco (vendas@genone.com.br)
Peptídeos catalogados, pedido mínimo de 2mg .
Tabela em breve
Entre em contato conosco (vendas@genone.com.br)
Principais bancos de dados mundiais de peptídeos bioativos.
Tabela em breve
Entre em contato conosco (vendas@genone.com.br)
A GenOne Biotechnologies propõe a seguir uma tabela com a pureza dos peptídeos recomendada para os ensaios mais comuns solicitados pelos nossos clientes.
Pureza recomendada
Aplicações
Imunologia Pureza - 70%
- Antígeno para a produção e purificação por afinidade de anticorpos policlonais
- Testes de ELISA
- Peptide arrays
Bioquímica Pureza - 85%
- Mapeamento de epítopos
- Estudos semi-quantitativos de interação enzima-substrato
- Estudos de fosforilação
- Estudos de peptide blocking por Western blotting
- Ensaios in vitro
- Estudos de aderência celular
Alta Pureza Pureza - 95%
- Estudos quantitativos de interação receptor-ligante
- Ensaios quantitativos de inibição e bloqueio competitivos
- Estudos quantitativos de fosforilação
- Estudos quantitativosde proteólise
- Estudos de NMR
- Estudos in vivo
- Geração de curvas de padrão
Industrial Pureza - 98%
- Peptídeos BPF para estudos de drogas
- Peptídeos para a indústria cosmética
- Ensaios clínicos
- Estudos de relação estrutura x atividade (SAR)
Observações:
Quando um peptídeo é entregue com pureza de 95%, isso significa que 95% do conteúdo de peptídeo entregue é composto pelo peptídeo solicitado (o que difere do conteúdo total do peptídeo). Os outros 5% de material peptídico é usualmente composto do que é chamado de “sequências deletadas” que usualmente podem ser co-purificadas com o peptídeo desejado durante a HPLC.
As sequências deletadas são geradas durante a síntese dos peptídeos, quando devido a ineficiência da reação de acoplamento, alguns aminoácidos são omitidos das moléculas sintetizadas. A pureza é determinada por HPLC de fase reversa.
É importante entender a diferença entre conteúdo peptídico e conteúdo total de peptídeos. Os peptídeos secos e entregues na forma de pó, normalmente não possuem somente peptídeos, mas também algumas substâncias como água, solventes adsorvidos, contra-íons e sais. Devido a própria natureza dos peptídeos, dificuldade de solubilidade, haverá quase sempre outros aditivos de forma a permitir solubilidade e estabilidade.
O conteúdo total se refere a esta mistura enquanto o conteúdo peptídico refere-se à fração do seu peptídeo na mistura. O conteúdo peptídico é usualmente 50-80% da massa total de pó enviado, esse conteúdo só é obtido precisamente pela análise do conteúdo de aminoácidos ou quantidade de nitrogênio que pode ser solicitada como um serviço à parte, bem como a remoção de TFA.
Existe uma maneira de se estimar o conteúdo peptídico teórico, assumindo que o único conteúdo não peptídico presente na amostra são os contra-íons. É possível fazer isso dividindo a massa molecular do peptídeo pela soma da massa molecular com o número de contra-íons necessários para neutralizar o peptídeo. O número de contra-íons é multiplicado pela massa molecular do TFA (MW=114).
Por exemplo, um peptídeo com massa molecular de 1500, com N-Terminal livre e uma Arginina, o seu conteúdo líquido é = 1500/(1500+2×114)= 0,87 ou 87%. Na prática, os contra-íons não são os únicos contaminantes, como citado anteriormente há também algumas substâncias como água, solventes adsorvidos e sais.
A seguir mostramos a grande diversidade de modificações que podem ser realizadas ao fazer o seu pedido com a GenOne Biotechnologies, porém caso deseje alguma que não esteja na lista não hesite em solicitar mais informações.
Modificações N-Terminal
Modificações C-Terminal
Modificações diversas
Acetilatação (Ac)
Amidação
Biotinilação, Fluoróforos
Formilação (For)
AMC
Pontes dissulfeto
Ácidos Graxos
Aldeído
Fosforilação
Benzoil (Bz)
Alcool
Conjugação com BSA, KLH, OVA
Benziloxicarbonilação (CBZ)
Éster (OMe, OEt)
PEGlação
Bromoacetil (Br-Ac)
p-Nitroanilina (pNA)
MAPS
Piroglutamil (pGlu) (Pyr)
NHMe, NHEt e NHisopen
Peptídeos Cíclicos
Succinilação (Suc)
tBu
Fluorescência e Quenchers
Tert-Butoxicarbonil (Boc)
TBzl
Aminoácidos Especiais
3-Mercaptopropil (Mpa)
Cisteamida (Cya)
Metil e N-Metil
Os CPPs são capazes de transportar ácidos nucleicos e nanomateriais para o interior das células.
Tabela em breve
Entre em contato conosco (vendas@genone.com.br)
Peptídeos catalogados, pedido mínimo de 2mg .
Tabela em breve
Entre em contato conosco (vendas@genone.com.br)
Principais bancos de dados mundiais de peptídeos bioativos.
Tabela em breve
Entre em contato conosco (vendas@genone.com.br)
Fale conosco
Estamos ansiosos para atender
às suas necessidades.
- (21) 3285-9105
- vendas@genone.com.br